Archiving
This page describes the archiving features in version 3.2 or later. If you're using version 3.0 or 3.1, please read the version 3.0/3.1 page.
With the archiving feature, you can create archive files that store all data published by your BioCASe web service. That might be handy if your database stores a huge number of records (several hundred thousands or even millions) and the traditional harvesting approach of paging through the records published by the web service takes unacceptably long. Once stored in an archive, this file can be downloaded from your web server or sent to the harvester via email and ingested much faster.
There are two types of archives: XML Archives simply store the records as compressed XML documents, each holding a customizable number of records. If your web service publishes 500,000 records by using the ABCD data schema, for example, an ABCD Archive created for that service could hold 500 ABCD documents, each storing 1,000 records. For each schema supported by a BioCASe web service, a separate XML Archive can be created. You can download a sample ABCD Archive here: File:AlgenEngelsSmall ABCD 2.06.zip.
DarwinCore Archives (DwCAs), in contrast, consist of one or several text files storing the core records and related information, zipped up together with a descriptor file and a metadata document. Similar to a BioCASe web service, DwCAs can be used to publish data to the Global Biodiversity Information Facility (GBIF). A detailed specification can be found on the TDWG site, a sample archive here: File:Desmidiaceae Engels.zip.
The BioCASe Provider Software can create XML archives for each schema supported by a web service. Information published by web services using the the ABCD 2.06 or HISPID 5 schema (dubbed as “ABCD/HISPID dumps”) can also be stored in one or several DarwinCore Archives. This is due to the fact that one BioCASe web service can publish several datasets, and a DarwinCore Archive can only store a single dataset. Unsurprisingly, DwCAs make use of the DarwinCore data standard, which is flat and less complex than ABCD. Consequently, a DarwinCore Archive will store less information than an ABCD dump for a given web service.
Contents
XML Archives
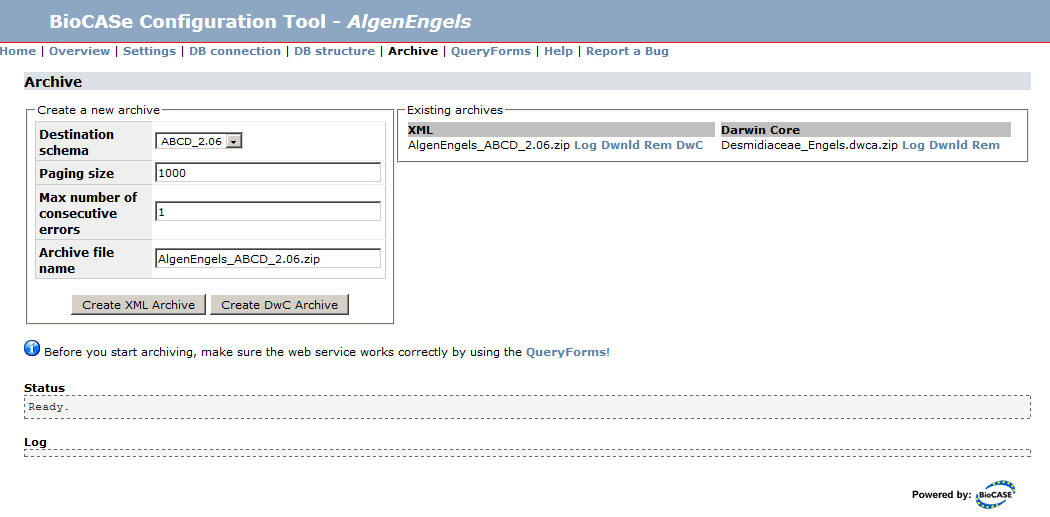
Once you’ve finished mapping a schema (see ABCD2Mapping to learn how to map ABCD, for example) and tested the resulting web service, you can create an XML Archive for that web service. To go to the archiving page, simply use the Archive link on the overview page of the datasource setup. If you haven’t mapped a schema, you won’t be able to create any archives. Before archiving, you should always make sure the web service returns correct results by using the QueryForms.
Creating an Archive
In the box “Existing Archives” you will see a list of already existing XML and DwC archives. If you go to the archiving page for the first time, it will be empty. Once you’ve created archives, they will show up here and can be downloaded and removed by using the links next to the entries.
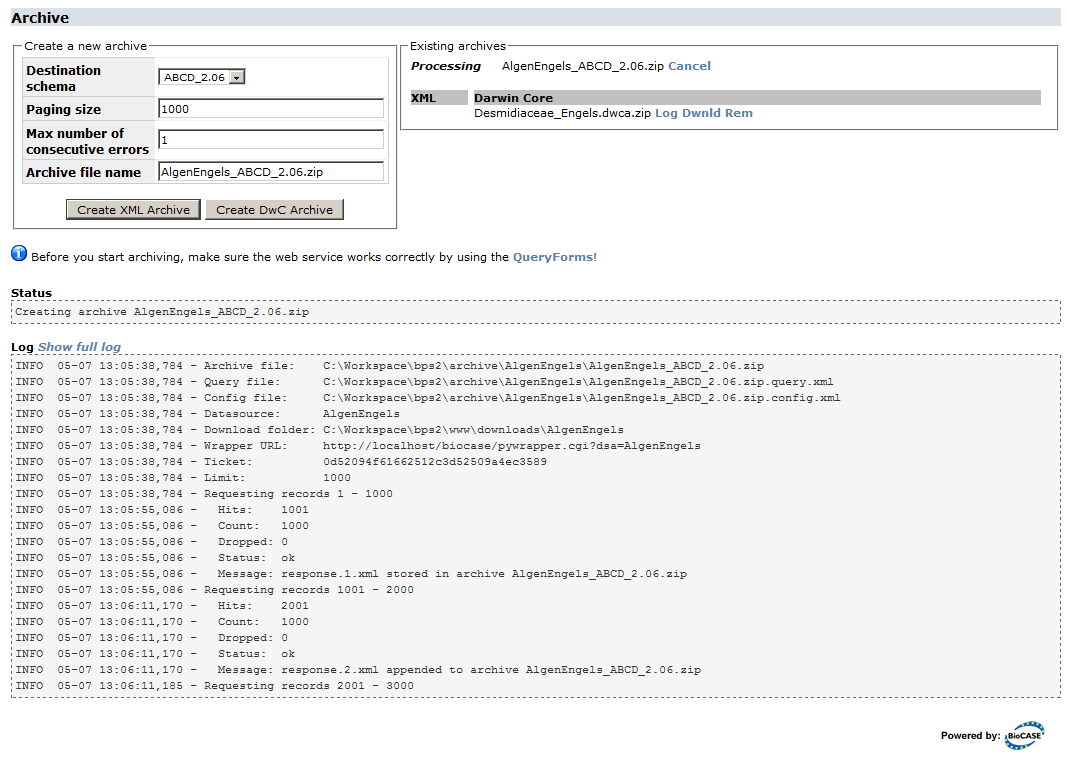
Create XML Archive allows you to create a new XML archive for each schema mapped. Simply select the schema and press Create Archive (leave the defaults unless you have a good reason to change them). During the archiving process, the Log will display messages indicating the progress:
You can always cancel the archiving process by pressing Cancel next to the name of the archive being currently processed. However, it will take some time for the cancelling to show effect, namely until the current paging step has been completed. Depending upon the paging size set and the speed of your server, this can be up to one minute.
As the archiving progresses, the first lines of the log will drop out of view. Pressing Show full log will open the full log in a separate browser tab. You can navigate away from the archiving page during the process (or simply close it) and return later. Once finished, the dump will show up in the XML list, or, if it’s still being processed, as processing. The links Log, Dwnld and Rem next to an existing archive will display the log, download or remove the file from the server. If the process failed for some reason (or you decided to cancel it), a new section Logs of aborted/failed runs will be displayed with a link for displaying the log file. This will allow you to find out what caused the abortion.
For a given web service, you can only create one archive at a time. Even though you can start archiving for several web services of your BioCASe installation in parallel, it is not a good idea. The archiving process will put heavy load on your servers – both the database server for data retrieval and the BioCASe server for creating the XML documents. So it is advisable to create just one archive at a time in order to allow both servers to respond to other requests.
Customizing the Archive
There are four parameters for customizing the archiving process:
Destination schema: Data schema that will be used for storing the published records in the archive.
Paging size: Number of records stored in each document. The default 1,000 should be suitable for most cases. Be aware that a BioCASe web service can restrict the number of records per request - if that limit is below the paging size set for archiving, the paging size will be adjusted automatically after the first packet of records was received from the web service.
Max number of consecutive errors: The default for this is “1”. That means that if an error occurs during data retrieval or XML/archive creation, the archiving process will be aborted. If you set this to a value larger than 1, the process will be cancelled after this number of errors. This allows you to complete the archiving in spite of errors, view the log file afterwards and have a look at the problematic records. Once you’ve corrected all errors breaking the dump, you should set the threshold to 1 and redo the archiving to create an archive storing all records.
Archive file name: Name of the archive file to be created. The default will be constructed by concatenating web service name and data schema.
DarwinCore Archives
For web services that use the ABCD 2.06 or HISPID 5 schema, all information published by the web service can be stored in DarwinCore Archives. The DarwinCore data standard is less complex than ABCD, so in most cases the DarwinCore Archive will not store all information published by the web service. Since one BioCASe web service can publish several datasets and a DarwinCore Archive can only store a single dataset, DwC-archiving a web service might result in several DarwinCore Archives.
DarwinCore archives are not created directly, but from the respective ABCD archive. So DwC-archiving a web service will usually first create an ACBD archive, which will then be converted into the respective DwC archive (alternatively, you can choose to convert an already existing ABCD archive into DwC).
Preparation
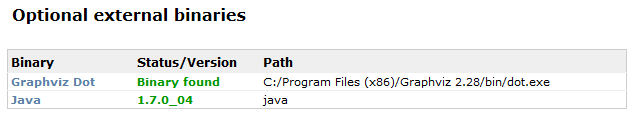
The conversion of ABCD into DwC is done through the Java-based Kettle library. Therefore, you need to have a Java Runtime Environment (version 1.5 or later) installed. You can check this on the libary test page under Optional external binaries:
If you don't have one, you can get it here. Important: If you're running BioCASe on a 64bit machine, make sure to get a 64bit Java, since this will boost performance considerably.
Once Java is installed, go to the System Administration (Start > Config Tool > System Administration) to check and adapt the conversion settings to your environment:
Java binary: If you're sure Java is installed, but the test lib page shows not installed, please provide the full path to the Java binary (for example /usr/local/jdk1.6.0/bin/java). If you have several Java versions installed (for example 32bit and 64bit versions), make sure to point to the correct (preferrably 64bit) version.
Max memory usage for Java VM (MB): This is the maximum amount of heap memory Java will be allowed to use (in MB). The default of 1024 will be sufficient for small datasets (up to 100,000 records). For medium-sized datasets (up to 1m records), you should set this value 2048. For large datasets with millions of records, a value of 4096 is recommended. If your server has enough free memory, you should be generous with this limit - larger values speed up the transformation process on most machines. However, you should make sure to leave enough memory for other required applications on your server.
Sort buffer size for transformation (number of rows): This is the number of rows kept in memory during sorting steps. The default of 100,000 should be OK for most purposes. If you run into memory problems and cannot increase the memory usage, you can lower this value to 50,000 or even 10,000 (do not enter thousands separators into the box). Even lower values will result in heavy disk usage and poor performance.
Creating an Archive
blubb
Output
The DwCA(s) created can be found in the folder output created in the base directory (dwca). Remember that there will be one DwCA per dataset. So depending upon how many datasets are stored in the source ABCD archive, there will be one or several files in the output folder.
On Linux and MacOS, the shell script dwca.sh will zip all files that make up one DarwinCore Archive into an archive file named after the dataset title. So you could send this files directly to a potential DwCA consumer:
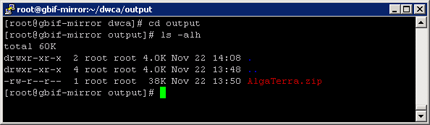
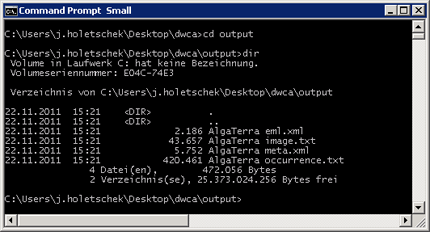
On Windows, the batch file dwca.bat of the prototype doesn't do that zipping. So if you go to the output file after the conversion has finished, you will see several text files and XML documents, all named after the pattern <dataset> <filename>. So for the example above, the output folder would look like this:
In order to create a DwCA that can be sent to a consumer, please do the follwing for each <dataset>:
- Note down the dataset title,
- Remove the dataset title in the file names (<dataset> <filename> becomes <filename>),
- Create a zip named <dataset>.zip with all the files you just renamed.
This lack of functionality is due to the limited capabilities of Windows batch files. Once integrated into the Provider Software, you won't need to zip up files manually.
Trouble Shooting
If you run into problems when using this tool, please contact us for help.
The default settings of the transormation should be working for small and medium-sized collections (up to 500,000 records). If the processing is cancelled with an OutOfMemoryError, you should try the follwing:
Increasing the maximum memory used by the transformer
Per default, the transformer will use up to 1GB of RAM for the transformation process. In case of an OutOfMemoryError, you should increase this limit to 2GB. Don't be stingy with memory, since it will be only used during the tranformation process and be freed afterwards. Moreover, this is only the upper limit, so actual memory usage might well stay below this limit.
To allow the transformer to use more memory, go to the folder kettle of your installation. Open the file kitchen.sh (Linux/MacOS) or kitchen.bat (Windows) in a text editor. Use the Find feature of the editor to go to the spot in file that specifies the memory limit. Replace the value 1024 with the amount of memory you want to allow the transformation to use, so e.g. 2048 for 2GB:
Save the file and restart the transformation.
Lowering the number of records stored in memory
Per default, Kettle will store up to 100,000 records in memory before starting to use temporary files for sorting rows. You can set this threshold to a lower value to save some memory. However, be advised that this will result in heavy disk I/O on your machine, slowing down the transformation. Therefore you should first increase the memory limit before changing this setting.
If you decide to do this, just add the desired maximim number of records to be stored in memory as a second parameter when invoking the transformer. So on Linux, type in
./dwca.sh AlgaTerra_ABCD_2.06.zip 10000
in order to set the threshold to 10,000 records. On a Windows machine, type in following:
dwca.bat ..\AlgaTerra_ABCD_2.06.zip 10000